Studies
SmallStruct has the tools to perform many types of structural studies on small molecules, or the interaction between small molecules and biological macromolecules. Below is a listing of the most common types of results the platform can deliver.
Structure elucidation/confirmation
What is in my test tube? The main service is to determine the chemical structure of unknown molecules that may be the result of anything from bio-prospecting to a chemical reaction with an unexpected outcome or a metabolite. This is achieved mostly by a combination of NMR and MS. MS is used to determine the exact mass from which the molecule formula that best fits the mass is calculated. MS fragmentation of the molecule can give additional information about substructures split from the mother ion. NMR is then used to determine the exact molecule framework, using various experiments correlating the through-bond scalar couplings and the through-space dipolar couplings. The sensitivity limit to do a full characterization is approximately 1 mM of compound in 120 µl solvent, corresponding to 50 µg @ 400 Da and preferably at least 1 mg of a peptide < 10 kDa. Smaller amounts can be analyzed to some extent, which may be enough if closely related analogues are already known or if MS sequencing is sufficient.
|
|
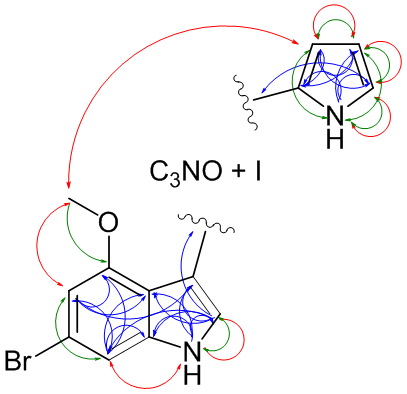 |
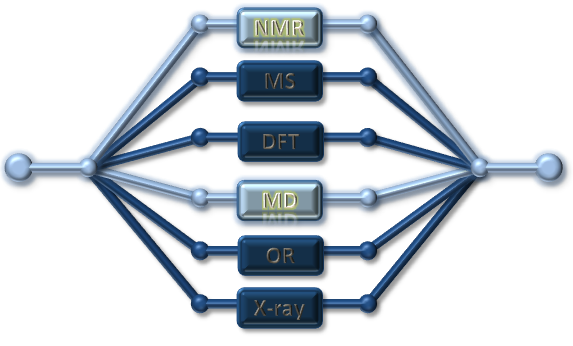
| Methodology involves NMR and MS, in certain cases DFT calculated couplings or chemical shifts are required | Schematic drawing of the NMR acquired correlations for Breitfussin A (red: noe, green: nJHH and blue: nJHC) |
Conformational analysis
Spectroscopic methods, mainly NMR, can together with computational methods be used to describe the conformation and distributions of conformation populations that small molecule adopts in solution, or in other media like for example a lipid bilayer. The dynamics of these processes can be monitored in many different time scales, ranging from fast nanosecond rotations to slow conformational exchange processes in the milliseconds to minutes time scale.
 |
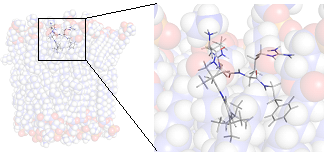 |
| Methodology involves NMR and MD in most cases | The Lytix pharmaceutical molecule LTX-109 modelled in a lipid bilayer |
Relative and absolute configuration
The determination of the absolute configuration is a challenging task that can be solved either by the direct approach of getting diffracting crystals, or by a combination of NMR, optical spectroscopies like ROA, VCD and/or ORD together with theoretical back-calculated spectra.
|
|
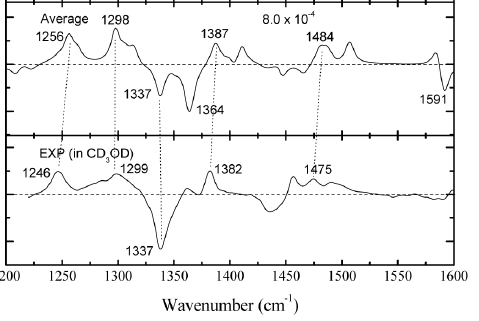 |
| Methodology involves either X-ray or a combination of NMR, DFT and one or more optical methods (and small amounts of MD) |
Simulated (top) and experimental (bottom) VCD spectra of R,R-Trp,Arg-diketopiperazine |
Interaction characterization
For macromolecules that are difficult to crystallize, the binding mode of a molecule binding to a macromolecule can be characterized by a NMR, ITC and computational methods taking advantage of experimental data. This can be either to just confirm or rule out binding by employing (competitive) fragment screening methods, or to determine the binding pose. ITC is used to describe the thermodynamics and stoichiometry of the binding process. The above can be done without any isotopic labeling. If 15N/13C double labeled protein can be expressed (requires decent expression yield in minimal media), an exact binding conformation can be determined at Ångstrom resolution given that the protein structure is known (by any structural method). Also the bound conformation of a flexible ligand can be determined if labeled protein is available.
 |
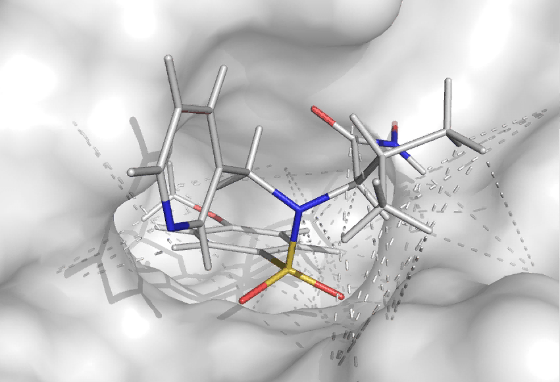 |
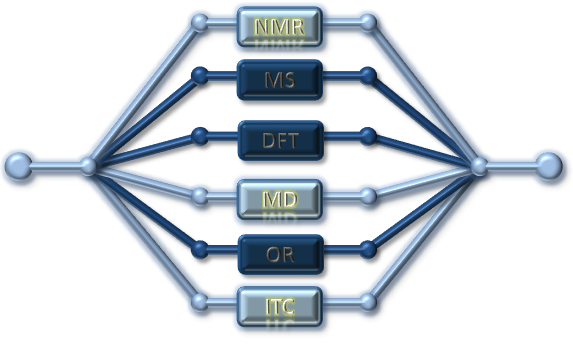
| Methodology involves NMR, ITC and MD | A small molecule docked in the active site of a target protein using NMR-constraints |
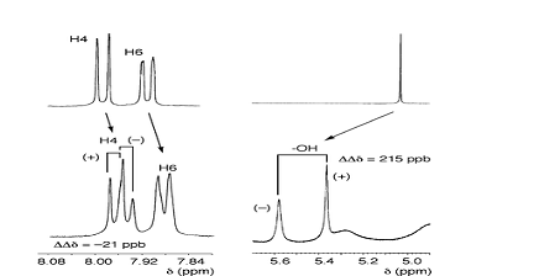
Enantiomeric purity/enantiomeric excess
The enantiomeric purity of a compound is most commonly determined by chiral HPLC, but we also have the possibility to set up NMR based methods using chiral solvent or derivatization to achieve enantiomer separation in cases when HPLC has difficulties.
 |
|
Associated platforms/centers
Academic collaborators
| Natural Products and Medicinal Chemistry, IFA, Tromsø | |
| Marine bioprospecting, Faculty of Biosciences, Fisheries and Economics, Tromsø | |
Commercial collaborators
Useful links